Genus of viruses
| Maricll/sandbox | |
|---|---|

|
|
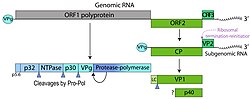
| Genome map of the genus Vesivirus (family Caliciviridae) showing the organization of ORFs encoding structural and non-structural proteins | |
| Scientific classification | |
| Kingdom: |
Orthornavirae |
| Phylum: |
Pisuviricota |
| Class: |
Pisoniviricetes |
| Order: |
Picornavirales |
| Family: |
Caliciviridae |
| Genus: |
Vesivirus |
| Synonyms | |
|
Rabbit calicivirus |
|
The Rabbit vesivirus (RaV) is a positive-sense, single-stranded RNA virus of the genus Vesivirus, family Caliciviridae. It was first isolated at the Veterinary Diagnostic Laboratory of Oregon State University from rabbits with gastrointestinal disease.[1] The virus is non-enveloped, isometric, and approximately 30 nanometers in diameter. Its genome is 8,295 nucleotides in length and contains three open reading frames (ORFs).[1] The genome encodes a 1,880-amino-acid polyprotein that undergoes proteolytic processing to yield the non-structural proteins required for replication. These include NS1-2 (unknown function), NS3 (NTPase/helicase), NS4 (membrane-associated protein involved in replication complex formation), NS5 (genome-linked viral protein VPg), and NS6/7 (a bifunctional protease and RNA-dependent RNA polymerase).[1]
RaV was classified within Caliciviridae based on its genome organization, proteolytic cleavage map, and phylogenetic and serological similarities to other vesiviruses.[1] While most caliciviruses cannot be propagated in vitro, Vesivirus species are notable for their ability to replicate in cell culture. Among them, RaV is unique in its capacity to reach high titers in a broad range of mammalian cell lines, including those of human origin such as Vero, RK13, 293T, HeLa, and HeLa-S3 cells.[1] This broad tropism distinguishes RaV from other vesiviruses, which typically exhibit strict host specificity.
Although RaV was originally identified in terrestrial rabbits, phylogenetic evidence suggests that it belongs to a subgroup of “marine-origin caliciviruses,” a lineage believed to have crossed into terrestrial hosts through contact or feeding on marine organisms.[1] Due to its close genetic relationship to the agent of vesicular exanthema of swine virus (VESV)—an eradicated, highly communicable disease—a moratorium on animal experimentation with RaV has been observed. As a result, knowledge about RaV’s natural host range and pathogenesis remains limited.[1]
A defining characteristic of RaV is its capacity to grow efficiently in diverse animal and human cell lines, making it an important model for studying calicivirus biology and replication.[1] This property also makes RaV a valuable tool for reverse genetics, a technique that enables recovery of infectious virus from cloned cDNA. In 2020, researchers reported the construction of a full-length RaV infectious cDNA clone under the control of a bacteriophage T7 RNA polymerase promoter.[1] Transfection of permissive cells with this plasmid, in the presence of T7 RNA polymerase supplied by a recombinant poxvirus helper, resulted in de novo synthesis of viral RNA and recovery of infectious RaV progeny. Infectious virus was also obtained when genome-length RNA transcripts were synthesized in vitro, capped, and subsequently transfected into cells.[1]
This reverse genetics system provides an efficient and reproducible means to study RaV replication, gene function, and pathogenesis. It also offers a platform for vaccine research, development of recombinant replicons, and antiviral screening, making RaV one of the most versatile experimental models among caliciviruses.[1]

Structure and genome
[edit]
Rabbit vesivirus (RaV) virions are small, non-enveloped particles typical of the Caliciviridae family. The viral genome is a positive-sense, single-stranded RNA molecule approximately 8,295 nucleotides in length, excluding a poly(A) tail of around 85 bases. The genome contains three open reading frames (ORFs) flanked by short untranslated regions (UTRs): a 19-nucleotide 5′ UTR and a 181-nucleotide 3′ UTR that is rich in uridines (44.2%).
ORF1 (nucleotides 20–5662) encodes a polyprotein of 1,880 amino acids containing conserved motifs characteristic of calicivirus nonstructural proteins. These include a helicase-NTPase (motif GxxGxGKT), a 3C-like cysteine protease (containing the catalytic cysteine motif), and an RNA-dependent RNA polymerase (motifs A and YGDD).
ORF2 (nucleotides 5668–7785) begins five nucleotides downstream of ORF1 and encodes a 705-amino-acid precursor of the major capsid protein (~78.7 kDa). Proteolytic processing produces the mature structural protein (~67 kDa).[2] Mass spectrometry of purified virions confirmed peptide sequences corresponding to the C-terminal region of this precursor, consistent with capsid maturation pathways described for Canine calicivirus and Feline calicivirus. ORF3 (nucleotides 7782–8114) overlaps ORF2 by four nucleotides and encodes a 110-amino-acid product, believed to function as a minor structural component of the virion, as observed in Rabbit hemorrhagic disease virus and Norovirus.[2] The 5′ UTRs of both genomic and subgenomic RNAs show a high degree of sequence conservation (12 of 22 nucleotides identical), suggesting a role in viral RNA replication, packaging, or transcriptional regulation.[2]
Phylogenetic analysis of the full ORF1 polyprotein shows that Rabbit vesivirus (RaV) clusters within the genus Vesivirus, grouping closely with Vesicular exanthema of swine virus (VESV), San Miguel sea lion virus (SMSV), Primate calicivirus Pan-1, and Walrus calicivirus (WCV).[2] Within Vesivirus, three major clades were Feline calicivirus isolates (FCV-F4, FCV-F9), RaV, Pan-1, and the marine vesiviruses (SMSV, VESV, WCV), and Canine calicivirus and isolate 2117 (unknown origin).[2] Sequence identity in the 5′ end of the genome (first 60 nucleotides) ranges from 46–98% among vesiviruses, contrasting with lower similarities to Lagovirus (10–13%) and Norovirus/Sapovirus (18–20%).[2]
Replication and Protein Processing
[edit]
RaV replication occurs in Vero (VMK) and PK-15 cells, producing cytopathic effects within 24–48 hours.[3] The viral RNA genome is linked at its 5′ end to a VPg protein, as demonstrated by loss of infectivity after proteinase K treatment.[3]
The ORF1 polyprotein undergoes proteolytic cleavage by its 3C-like cysteine protease.[3] Predicted cleavage sites occur at residues E435, Q791, E1070, and E1183, generating nonstructural proteins analogous to 2C (NTPase), 3A, 3B (VPg), and 3CD (protease-polymerase).[3] The 3CD product in RaV appears not to be further cleaved into separate 3C and 3D proteins, a trait shared with Feline calicivirus Pan-1, distinguishing vesiviruses from other calicivirus genera.[3]
RaV infects rabbits (Oryctolagus cuniculus), with intestinal epithelial cells being the primary target.[3] Initial studies in cell culture used PK-15 (porcine kidney) and Vero (African green monkey kidney) cell lines to propagate the virus, where it produced cytopathic effects (CPE) characterized by rounded, refringent cells detaching from monolayers.[3] Oral infectivity experiments in rabbits showed asymptomatic infection with seroconversion, suggesting limited tissue tropism in vivo following cell culture adaptation.[3]
Entry into the cell
[edit]
RaV enters host cells via an as-yet uncharacterized receptor-mediated mechanism.[4] In vitro, the virus successfully infects PK-15 and Vero cells, leading to cytopathic effects within 5–12 hours post-infection at a multiplicity of infection (MOI) of 1.[4] Like other caliciviruses, the genome contains a 5′ viral protein genome-linked (VPg) protein, essential for efficient translation and replication. Removal of VPg via proteinase K drastically reduces infectivity, highlighting its role in early post-entry processes.[4]
RaV exhibits a lytic replication cycle in permissive cells. Viral RNA released into the cytoplasm is translated into the ORF1 polyprotein, which is processed by the viral 3C-like protease into functional non-structural proteins, including the RNA-dependent RNA polymerase (3D) necessary for genome replication.[4] Subgenomic RNA is transcribed to produce structural proteins (ORF2 and ORF3), which assemble into virions.[4] Virions accumulate in the cytoplasm and are released upon cell lysis, producing clear plaques in vitro. Unlike some caliciviruses, the RaV 3CD polyprotein does not undergo further cleavage into separate protease and polymerase components, consistent with other Vesivirus members.[4]
Host Range and Pathogenicity
[edit]
RaV was first isolated from young New Zealand/California crossbreed rabbits (Oryctolagus cuniculus) exhibiting mild gastrointestinal disorders in Oregon (1995).[4] Experimental infection in rabbits using Vero cell-adapted RaV produced asymptomatic infections, characterized by seroconversion and limited replication in intestinal tissues, suggesting attenuation during cell culture adaptation.[4]
The discovery of RaV established that rabbits can host caliciviruses from multiple genera, including both Lagovirus (e.g., rabbit hemorrhagic disease virus) and Vesivirus. This demonstrates that genus naming within Caliciviridae is not strictly host-based.
Genome Organization
[edit]
| ORF | Position (nt) | Product | Description |
|---|---|---|---|
| ORF1 | 20–5662 | Non-structural polyprotein | ORF1 encodes a large non-structural polyprotein that is processed by viral proteases into multiple functional proteins (NS1–NS6/7).[5][6] These proteins are essential for viral replication, including enzymatic activities such as protease, helicase, and RNA-dependent RNA polymerase functions, which coordinate genome replication and processing of viral proteins.[5][6] |
| ORF2 | 5668–7785 | Major structural protein (VP1) | ORF2 encodes the major capsid protein VP1, which forms the bulk of the viral capsid and is critical for viral particle assembly, structural integrity, and host cell recognition.[5] VP1 is the primary antigenic determinant of the virus and is essential for infectivity.[5][6] |
| ORF3 | 7782–8114 | Putative minor structural protein (VP2) | ORF3 produces VP2, a smaller structural protein thought to play a supportive role in capsid assembly and stability.[5][6] VP2 may assist in proper folding of VP1 and contribute to the formation of infectious virions, though its exact functions remain under investigation.[6] |
| Misc | 8296–8380 | Poly-A tail | The miscellaneous region at the 3′ end of the genome contains the polyadenylated [poly(A)] tail, which is important for RNA stability, efficient translation, and replication.[6] The poly-A tail interacts with host cell machinery to facilitate viral protein synthesis.[5][6] |
Detection of RaV in cell culture and experimental systems involved a combination of molecular and immunological approaches designed to confirm both the presence of viral RNA and the expression of viral proteins.[5] Initially, total RNA was extracted from infected Vero cells and subjected to reverse transcription followed by polymerase chain reaction (RT-PCR) using primers specific for RaV genomic regions, including VP2, to detect viral genomes and any molecular tags introduced in engineered clones. RT-PCR served as a rapid and sensitive method to detect viral RNA in both cell lysates and culture supernatants, providing quantitative evidence of viral replication.[5] Controls without reverse transcriptase were included to rule out DNA contamination.
In parallel, viral protein expression was assessed by Western blot analysis. Protein lysates from infected or transfected cells were separated by SDS-PAGE, transferred to membranes, and probed with specific polyclonal antisera against RaV NS3 and VP1 proteins.[5] Detection of these viral antigens confirmed active viral replication and proper translation of structural and non-structural proteins. Secondary antibodies conjugated with infrared dyes allowed high-sensitivity visualization of protein bands.[5]
Additionally, the replication competence of rescued RaV progeny was evaluated through plaque assays in Vero cells. This method quantified infectious virus particles and enabled comparison of growth kinetics between wild-type RaV and engineered clones (e.g., rRaV/Xh).[5] Sequential passages in Vero cells ensured stable propagation, while blind passage experiments confirmed the infectivity of rescued viruses. One-step growth curves provided temporal resolution of viral replication, complementing molecular and protein-based detection.[5]
Together, these methods—RT-PCR, Western blotting, and plaque assays—offered a comprehensive and confirmatory approach to detect RaV, analogous to the layered strategies used for other viral systems such as EBV, where serological assays, nucleic acid amplification, and in situ hybridization are combined to determine infection status and stage.[5] This multi-modal detection strategy ensured high specificity in identifying RaV replication in experimental systems.
Rabbit vesivirus (RaV) infection induces extensive membrane remodeling and the accumulation of vesicular structures in the cytoplasm of infected Vero cells.[3] These alterations are associated with the formation of replication complexes (RCs)—membrane-bound structures that serve as the site of viral RNA synthesis.[2] Within these virus factories, viral nonstructural proteins (2AB-, 2C-, 3A-, 3B-, and 3CD-like), viral RNA, and the endoplasmic reticulum (ER)-resident protein calnexin co-localize, indicating that replication occurs in association with modified ER-derived membranes.[2][5]
RaV RCs protect their endogenous RNA templates from nuclease degradation, rendering them inaccessible to exogenous synthetic RNA.[5] These structures are, however, permeable to exogenously added recombinant VPg, which can be nucleotidylylated within the RCs, though no RNA chain elongation from either native or recombinant VPg has been demonstrated in vitro.[5]
Quantitative RT-PCR analyses revealed that subgenomic RNA (sgRNA) is more abundant than genomic RNA (gRNA) in RaV RCs, suggesting that sgRNA synthesis is an early replication event that supports efficient capsid protein production.[5] The overrepresentation of sgRNA could also explain reports of its packaging into lower-density virions, as seen in related caliciviruses such as Feline calicivirus (FCV) and Rabbit hemorrhagic disease virus (RHDV).[5]
The replication complexes are intimately linked to host cell membranes, particularly the ER, as shown by the co-fractionation of viral nonstructural proteins and calnexin in high-density sucrose gradients.[5] Detergent treatment disrupts this association, leading to loss of polymerase activity, highlighting the importance of membrane integrity for replication.[2]
The viral proteins present in purified RCs include mature forms of 2C, 3A, and 3B (VPg), as well as the major capsid protein. The bifunctional 3CD-like protein and the 2AB product remain unprocessed in RaV, a notable feature distinguishing Vesivirus replication from that of other Caliciviridae genera.[5] The RCs also contain both RNA-linked and free VPg, the latter capable of undergoing uridylylation in vitro.[3] However, no evidence supports RNA chain elongation from VPg, leaving unresolved whether VPg acts as a true primer or as a capping protein attached to viral RNA during later stages of replication.[2][5]
Immunofluorescence microscopy shows that RaV nonstructural proteins localize to discrete cytoplasmic and perinuclear foci, overlapping with ER markers, consistent with the formation of membrane-associated multiprotein replication complexes.[3][5] These structures are analogous to the replication sites described for Hepatitis C virus and other positive-strand RNA viruses.[2]
Rabbit vesivirus (RaV) was first isolated from New Zealand/California crossbred rabbits (Oryctolagus cuniculus) showing mild gastrointestinal symptoms in Oregon in the mid-1990s.[5] The virus was later characterized as a new member of the genus Vesivirus within the family Caliciviridae, based on genomic organization and sequence similarity to San Miguel sea lion virus and vesicular exanthema of swine virus.[5]
The full-length RaV genome (~8.3 kb) was sequenced and found to contain three open reading frames (ORFs) typical of Caliciviridae: ORF1 encoding the nonstructural polyprotein, ORF2 encoding the capsid precursor, and ORF3 encoding a small structural protein.[5] This genomic structure and the presence of a bifunctional 3CD-like protease–polymerase supported its classification as a vesivirus distinct from Lagovirus and Norovirus species infecting rabbits.[2]
The study of Rabbit vesivirus has contributed to understanding the replication strategies of caliciviruses, particularly their use of membrane-bound replication complexes. RaV provides a valuable experimental model for examining viral–host interactions because it can be cultured in Vero cells and yields active replication complexes in vitro.[5]
These RCs maintain enzymatic activity capable of incorporating nucleotides into RNA products resembling viral gRNA and sgRNA.[3][5] Although the exact mechanism of RNA initiation remains unclear, these in vitro systems preserve native protein–protein and protein–membrane interactions, making them superior to simplified recombinant-protein assays previously used in calicivirus studies.[2]
The RaV system has also been proposed as a platform for investigating antiviral compounds that target calicivirus replication. Its membrane-associated RCs offer a physiologically relevant environment for testing inhibitors of RNA polymerase or VPg uridylylation activity.[1][2][3][5]
- Álvarez ÁL, Garcia-Manso A, Dalton KP, Martin-Alonso JM, Nicieza I, Podadera A, Acosta-Zaldívar M, de Llano D, Parra F (2020). “Reverse Genetics System for Rabbit vesivirus.” Frontiers in Microbiology. 11: 596245. doi:10.3389/fmicb.2020.596245. PMC 7693663. PMID 33304341.
- Casais R, Gonzalez Molleda L, Machin A, del Barrio G, Garcia Manso A, Dalton KP, Coto A, Martin Alonso JM, Prieto M, Parra F (2008). “Structural and functional analysis of virus factories purified from Rabbit vesivirus-infected Vero cells.” Virus Research. 137(1): 112–121. doi:10.1016/j.virusres.2008.06.009.
- Martin-Alonso JM, Skilling DE, Gonzalez-Molleda L, del Barrio G, Machin A, Keefer NK, Matson DO, Iversen PL, Smith AW, Parra F (2005). “Isolation and characterization of a new Vesivirus from rabbits.” Virology. 337(2): 373–383. doi:10.1016/j.virol.2005.04.018. PMID 15901487.
- Kwit E, Rzeżutka A (2019). “Molecular methods in detection and epidemiologic studies of rabbit and hare viruses: a review.” Journal of Veterinary Diagnostic Investigation. 31(4): 497–508. doi:10.1177/1040638719852374.
- Salomé González-Reyes, García-Manso A, del Barrio G, Dalton KP, González-Molleda L, Arrojo-Fernández J, Nicieza I, Parra F (2009). “Role of annexin A2 in cellular entry of rabbit vesivirus.” Journal of General Virology. 90(11): 2654–2662. doi:10.1099/vir.0.013276-0.
- Parra F (2004). “Rabbit vesivirus complete virion genome.” GenBank: AJ866991.1. Available at: https://www.ncbi.nlm.nih.gov/nuccore/AJ866991
- Martin-Alonso JM, Skilling DE, González-Molleda L, del Barrio G, Machin Á, Keefer NK, Matson DO, Iversen PL, Smith AW, Parra F (2005). “Isolation and characterization of a new Vesivirus from rabbits.” Virology. 337(2): 373–383. https://doi.org/10.1016/j.virol.2005.04.018
- ^ a b c d e f g h i j k l Álvarez, Ángel L.; García-Manso, Alberto; Dalton, Kevin P.; Martín-Alonso, José M.; Nicieza, Inés; Podadera, Ana; Acosta-Zaldívar, Maikel; de Llano, Daniel; Parra, Francisco (2020-11-13). “Reverse Genetics System for Rabbit vesivirus”. Frontiers in Microbiology. 11. doi:10.3389/fmicb.2020.596245. ISSN 1664-302X. PMC 7693663. PMID 33304341.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ a b c d e f g h i j k l m n “Structural and functional analysis of virus factories purified from Rabbit vesivirus-infected Vero cells”. Virus Research. 137 (1): 112–121. 2008-10. doi:10.1016/j.virusres.2008.06.009.
- ^ a b c d e f g h i j k l m Martín-Alonso, José M.; Skilling, Douglas E.; González-Molleda, Lorenzo; del Barrio, Gloria; Machín, Ángeles; Keefer, Nathan K.; Matson, David O.; Iversen, Patrick L.; Smith, Alvin W.; Parra, Francisco (2005-07). “Isolation and characterization of a new Vesivirus from rabbits”. Virology. 337 (2): 373–383. doi:10.1016/j.virol.2005.04.018.
- ^ a b c d e f g h Kwit, Ewa; Rzeżutka, Artur (2019-07). “Molecular methods in detection and epidemiologic studies of rabbit and hare viruses: a review”. Journal of Veterinary Diagnostic Investigation. 31 (4): 497–508. doi:10.1177/1040638719852374. ISSN 1040-6387. PMC 6857032. PMID 31131728.
- ^ a b c d e f g h i j k l m n o p q r s t u v w x y z aa ab González-Reyes, Salomé; García-Manso, Alberto; del Barrio, Gloria; Dalton, Kevin P.; González-Molleda, Lorenzo; Arrojo-Fernández, José; Nicieza, Inés; Parra, Francisco (2009-11-01). “Role of annexin A2 in cellular entry of rabbit vesivirus”. Journal of General Virology. 90 (11): 2724–2730. doi:10.1099/vir.0.013276-0. ISSN 0022-1317.
- ^ a b c d e f g “Rabbit vesivirus complete virion genome”. 2005-06-20.