XK Related 7 (XKR7) is a protein predicted to be a transmembrane protein in Homo sapiens encoded by the XKR7 gene. The primary alias is c20orf159, but it is commonly referred to as XKR7. It is predicted to be involved in apoptotic process during development.[1][2]
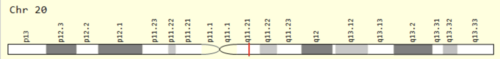
The XKR7 gene is located on plus end strand of chromosome 20 at locus 20q.11.21 and spans from 31968151 to 32003387.[1] It is 1740 base pairs long and contains 3 exons with no known isoforms.[1][3]

Several other genes are near the XKR7 gene along chromosome 20. Some of the closest ones are PDRG1, TTLL9, and CCM2L.[3]
| Gene | Description |
|---|---|
| PDRG1 | A protein-coding gene near XKR7 in the regulatory-element context of its promoter/enhancer region. |
| TTLL9 | Also appears in the same regulatory neighborhood as XKR7. |
| CCM2L | Another gene near XKR7 whose protein interacts with XKR7 |
Gene Level Regulation
[edit]
XKR7 is expressed in the brain and pancreas, but lowly at around fifteen percent.[4] The multiple sequence alignment of XKR7 and it’s orthologs showed high conservation of the 5′ untranslated region where transcription factor ZNF460 binds too.
The XKR7 protein contains 579 amino acids with a molecular weight of 63 kDa, and consists of 3 disordered regions, alongside 6 transmembrane regions.[5][1] XKR7 was found to have high amounts of these amino acids: Leu,Val, Ile, Phe, and Met; all of which contribute to transmembrane helices.[6] There are no major charge clusters, showing no strong cytoplasmic regulatory domains. However, the isoelectric point is 9.2, so at lower pHs, XKR7 will be positively charged.[6]

The tertiary structure of XKR7 mostly consists of alpha helices. The most confident parts of the structure is the transmembrane region, which is highly hydrophobic and uncharged. The disordered regions are shown by red and blue, which consists of cations and anions, and is hydrophilic.
Protein Level Regulation
[edit]
There are around 7 phosphorylation sites between 2nd and 3rd transmembrane protein. Analysis on the mouse ortholog produces the exact same image with these phosphorylation sites conserved: S154, G167, T183, Y201, S206, S240, and S249.[7] Y201 is the most functionally important site. When Y201 is phosphorylated, it’s a binding site for SH2-domain proteins (docking site), and when it’s unphosphorylated it functions as a tyrosine-based sorting signal for adaptor proteins.
Protein – Protein Interactions
[edit]

CCM2L and PGBD5 have the highest interaction scores. Using co-IP or pull-down assays could test physical binding. Using fluorescence microscopy would verify whether the proteins are in the same cellular compartment.
| Abbreviated name | Full Name | Score | Subcellular Compartment | Function |
| CCM2L | Cerebral cavernous malformations 2 protein-like | 0.628 | Membrane associated | Regulates signaling and cell-cell junctions |
| PGBD5 | PiggyBac transposable element-derived protein 5 | 0.608 | Nucleus | Influences genome regulation in neurons |
| CCDC137 | Coiled-coil domain containing 137 | 0.529 | Nucleus/Nucleolus | Nucleolar organization |
| ZC3H3 | Zinc finger CCCH domain-containing protein 3 | 0.501 | Nucleus | Regulates RNA processing |
Single Nucleotide Polymorphisms
[edit]
Most of the SNPs were found towards the C-term, transmembrane regions, or disordered regions.
| Order in CT | SNP | Change | Base Change | Mutation Type | Significance |
| 1 | rs138804760 | Met417Thr | T → C | missense | TMR 7 |
| 2 | rs147437006 | Arg292Glu
Arg292Leu |
G → A,T | missense | N/A |
| 3 | rs200124372 | Arg483Glu
Arg483Pro |
G → A,C,T | missense | In disordered region |
| 4 | rs367920055 | Thr558Lys
Thr558Met |
C → A,T | missense | C-term |
| 5 | rs371260952 | Asp374Lys | C → A,T | missense or silent | TMR 6 |
| 6 | rs541196906 | Arg565Glu
Arg565Pro |
G → A,C,T | missense | C-term |
| 7 | rs546872871 | Ala161Met
Ala161Val |
C → A,G,T | missense | In disordered region |
| 8 | rs572256162 | Val233Tyr | G → A | missense | C-term |
| Paralog | Amino Acid Lenght | Percent Similarity | Percent Identity |
|---|---|---|---|
| HSA_XKR4 | 650 | 57% | 46% |
| HSA_XKR6 | 641 | 56.3% | 44.60% |
| HSA_XKR8 | 395 | 33.10% | 22.30% |
| HSA_XKR9_2 | 373 | 30.70% | 18.10% |
| HSA_XKR5_1 | 686 | 26.40% | 18.10% |
| HSA_XKR5_2 | 523 | 16.20% | 11.00% |
| HSA_XKR9_1 | 241 | 19.80% | 12.80% |
XKR7 in Homo Sapiens has many paralogs as it’s a part of a very large X-Kell related protein family. The XKR protein family primarily functions as lipid scramblases, a crucial protein to signal for apoptosis.[8] There are a total of seven paralogs with the two most similar to XKR7 are XKR4 and XKR6 (Ref. Paralog table). Paralogs were found using Genecards and NCBI Protein database, while the sequence similarity and identity were done through Emboss Needle.[9][3][1]
| Genus and species | Common Name | Taxanomic Group | Median Date of Divergence (MYA) | Accession Number | Amino Acid Sequence Lenght | Sequence Identity to Human Protein | Sequence Similarity to Human Protein |
| Homo sapiens | Humans | Primates | 0 | NP_001011718.1 | 579 | 100% | 100% |
| Sarcophilus harrisii | Tasmanian Devil | Dasyuromorphia | 160 | XP_023363111 | 571 | 72% | 78.10% |
| Gracilinanus agilis | Agile Gracile Opossum | Didelphimorphia | 160 | XP_044515717 | 625 | 76% | 75.00% |
| Opisthocomus hoazin | Hoazin | Opisthocomiformes | 319 | XP_075294519 | 545 | 68% | 75.40% |
| Excalfactoria chinensis | King Quail | Galliformes | 319 | XP_072206629 | 527 | 68% | 75% |
| Malaclemys terrapin pileata | Diamondback Terrapin | Testudines | 319 | XM_054046095.1 | 577 | 70% | 79.60% |
| Podarcis raffonei | Aeolin Wall Lizard | Squamata | 319 | XP_053247703 | 574 | 72% | 75.60% |
| Erythrolamprus reginae | Royal Ground Snake | Squamata | 319 | XP_070600370 | 590 | 66.70% | 75.60% |
| Chelonoidis abingdonii | Pinta Island Tortoise | Testudines | 319 | XP_074928543 | 581 | 67.30% | 75.00% |
| Pleurodeles waltl | Iberian Ribbed Newt | Urodela | 352 | XP_069099477 | 531 | 66% | 74.70% |
| Microcaecilia unicolor | Microcaecilia unicolor | Gymnophiona | 352 | XP_030069193.1 | 562 | 64% | 72.40% |
| Phyllobates terribilis | Golden Poison Frog | Dendrobatidae | 352 | XP_073538933 | 494 | 62% | 67.70% |
| Conger conger | European conger | Anguilliformes | 429 | XP_061114556 | 567 | 63% | 73.40% |
| Polypterus senegalus | Gray bichir | Polypteriformes | 429 | XP_039590706 | 554 | 63% | 72.10% |
| Erpetoichthys calabaricus | Reedfish | Polypteriformes | 429 | XP_028667497 | 554 | 63% | 71.90% |
| Semicossyphus pulcher | California Sheephead | Perciformes | 429 | XP_069515596 | 611 | 56% | 68.30% |
| Callorhinchus milii | Australian Ghostshark | Holocephali | 462 | XP_042197801 | 549 | 61% | 68.30% |
| Chiloscyllium punctatum | Brownbanded Bamboo Shark | Orectolobiformes | 462 | XP_072412862 | 595 | 60% | 71.60% |
| Hemitrygon akajei | Red Stingray | Myliobatiformes | 462 | XP_072917960 | 574 | 59% | 70.80% |
| Amblyraja radiata | Thorny Skate | Rajiformes | 462 | XP_032897124 | 581 | 64% | 70.30% |
| Petromyzon marinus | Sea Lampray | Petromyzontiformes | 563 | XP_075929753 | 752 | 53% | 47.60% |
Orthologs of XKR7 Homo sapiens are sorted by median date of divergence and then by sequence identity to XKR7 Homo sapiens. There were an overwhelming number of mammal, avian, and reptilian orthologs, but the number of orthologs decreased as the median date of divergence increased. Actinopterygii and Chondrichthyes both only had a handful of orthologs for XKR7 Homo sapiens, but it mostly consisted of orthologs for XKR7 Homo sapiens paralogs. Sequences, identity, and similarity were found using NCBI Protein and Emboss Needle. [9]Cyclostomata, Jawless fish, are the most distantly related species to humans with the Sea Lamprey having a median date of divergent of 563 million years ago and 53% identity. Marsupials, Avians, Reptilia, with average sequence identity of 73.6% sequence identity.
Clinical Significance
[edit]
It is suggested, using mouse shotgun data alongside humans, could hold the key to understanding diseases like Creutzfeldt-Jakob disease, a rare and fatal neurodegenerative disease from an accumulation of prions, and severe immunodeficiency.[10][11] A study by Zhang et al. (2020) used RNA-seq for comparative expression in small cell lung cancer (SCLC) and adjacent normal tissue, which identified DRD2 and XKR7 upregulating SCLC.[12] Such a significant difference suggests that XKR7 could be linked to greater tumor aggressiveness. Gnathodiaphyseal Dysplasia is also associated with XKR7, but it’s not the main cause.[3]
- ^ a b c d e “XKR7 XK related 7 [Homo sapiens (human)] – Gene – NCBI”. www.ncbi.nlm.nih.gov. Retrieved 2025-09-25.
- ^ “XK-related protein 7 [Homo sapiens] – Protein – NCBI”. www.ncbi.nlm.nih.gov. Retrieved 2025-09-25.
- ^ a b c d GeneCards Human Gene Database. “XKR7 Gene – GeneCards | XKR7 Protein | XKR7 Antibody”. www.genecards.org. Archived from the original on 2025-08-15. Retrieved 2025-12-08.
- ^ “XKR7 XK related 7 [Homo sapiens (human)] – Gene – NCBI”. www.ncbi.nlm.nih.gov. Retrieved 2025-12-12.
- ^ “CCDS Report for Consensus CDS”. www.ncbi.nlm.nih.gov. Retrieved 2025-09-25.
- ^ a b EMBL-EBI; Institute, European Bioinformatics. “Job Dispatcher homepage | EMBL-EBI”. www.ebi.ac.uk. Retrieved 2025-12-12.
- ^ “ELM – unknown”. elm.eu.org. Retrieved 2025-12-12.
- ^ Suzuki, Jun; Imanishi, Eiichi; Nagata, Shigekazu (2014-10-31). “Exposure of Phosphatidylserine by Xk-related Protein Family Members during Apoptosis *”. Journal of Biological Chemistry. 289 (44): 30257–30267. doi:10.1074/jbc.M114.583419. ISSN 0021-9258. PMC 4215210. PMID 25231987.
- ^ a b “Job Dispatcher homepage | EMBL-EBI”. www.ebi.ac.uk. Retrieved 2025-10-20.
- ^ The Wellcome Trust Sanger Institute (December 2001). “The DNA sequence and comparative analysis of human chromosome 20”. Nature. 414 (6866): 865–871. Bibcode:2001Natur.414..865D. doi:10.1038/414865a. ISSN 0028-0836. PMID 11780052.
- ^ Zerr, Inga (2022-04-06). “Laboratory Diagnosis of Creutzfeldt–Jakob Disease”. New England Journal of Medicine. 386 (14): 1345–1350. doi:10.1056/NEJMra2119323. ISSN 0028-4793. PMID 35388668.
- ^ Zhang, J.; Ning, R.; Wu, Z.; Li, S.; Huang, J.; Lin, H.; Lin, W.; Chen, A.; Shi, S.; Chen, L.; Wu, L.; He, J. (May 2020). “RNA-seq Reveals DRD2 and XKR7 Were Upregulated in SCLC”. A70. Advances in Lung Cancer Therapeutics. American Thoracic Society. pp. A2480. doi:10.1164/ajrccm-conference.2020.201.1_MeetingAbstracts.A2480.