Protein-coding gene in the species Homo sapiens
Mitochondrial acyl carrier protein (mtACP), also known as NADH:ubiquinone oxidoreductase subunit AB1 (NDUFAB1), is a protein encoded by the human NDUFAB1 gene.[4] As a soluble matrix protein, it functions as a scaffold on which mitochondrial fatty acid synthesis (mtFAS) builds de novo fatty acyl chains.[5] The octanoyl (C8) form of mtACP provides the precursor for mitochondrial lipoic acid biosynthesis, and protein–protein interactions between mtACP and LYRM proteins are required for central mitochondrial processes, including the assembly of respiratory chain complexes, iron–sulfur cluster biogenesis, and mitochondrial ribosome assembly.[6] Through its binding to the complex I subunits LYRM3 and LYRM6, acyl-mtACP is incorporated as a structural component of complex I, giving rise to its designation as NDUFAB1.[7]
mtFAS-derived acyl-mtACP and its interactions with LYRM proteins have been hypothesized to form a feedback loop that allows acetyl-CoA to regulate its own consumption through the assembly of respiratory chain complexes, which in turn controls citric acid cycle flux.[8]
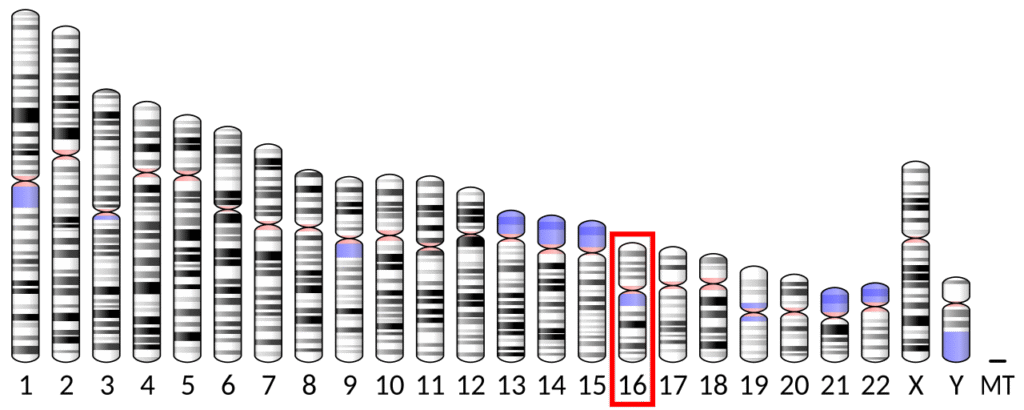
The NDUFAB1 gene is located on the p arm of chromosome 16 at position 12.2 and it spans 15,327 base pairs.[4] The NDUFAB1 gene produces a 17.4 kDa protein composed of 156 amino acids.[9][10] NDUFAB1 is a subunit of the enzyme NADH dehydrogenase (ubiquinone), the largest of the respiratory complexes. The structure is L-shaped with a long, hydrophobic transmembrane domain and a hydrophilic domain for the peripheral arm that includes all the known redox centers and the NADH binding site.[11] NDUFAB1 is one of about 31 hydrophobic subunits that form the transmembrane region of Complex I. It has been noted that the N-terminal hydrophobic domain has the potential to be folded into an alpha helix spanning the inner mitochondrial membrane with a C-terminal hydrophilic domain interacting with globular subunits of Complex I. The highly conserved two-domain structure suggests that this feature is critical for the protein function and that the hydrophobic domain acts as an anchor for the NADH dehydrogenase (ubiquinone) complex at the inner mitochondrial membrane.[4]
mtACP exists in apo, holo, and acylated forms with distinct biological functions:
The human NDUFAB1 gene codes for a subunit of Complex I of the respiratory chain, which transfers electrons from NADH to ubiquinone.[4] However, NDUFAB1 is an accessory subunit of the complex that is believed not to be involved in catalysis.[19] Initially, NADH binds to Complex I and transfers two electrons to the isoalloxazine ring of the flavin mononucleotide (FMN) prosthetic arm to form FMNH2. The electrons are transferred through a series of iron-sulfur (Fe-S) clusters in the prosthetic arm and finally to coenzyme Q10 (CoQ), which is reduced to ubiquinol (CoQH2). The flow of electrons changes the redox state of the protein, resulting in a conformational change and pK shift of the ionizable side chain, which pumps four hydrogen ions out of the mitochondrial matrix.[11]
Clinical Significance
[edit]
In a genome-wide association study (GWAS) meta-analysis conducted across European and African American populations, the NDUFAB1 gene and two other genes (MFAP3L and PALB2) were identified as genetic loci significantly associated with anxiety disorders (ADs). Since the comorbidity of ADs arises from their shared genetic basis, these candidate genetic loci may become therapeutic targets for AD treatments.[20] Moreover, a study to identify small molecule drug targets for Tetralogy of Fallot (TOF), a congenital malformation of the heart, found the NDUFAB1 gene to be a major hub gene of differentially expressed genes in TOF.[21]
- ^ a b c GRCh38: Ensembl release 89: ENSG00000004779 – Ensembl, May 2017
- ^ “Human PubMed Reference:”. National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ “Mouse PubMed Reference:”. National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b c d “Entrez Gene: NDUFAB1 NADH dehydrogenase (ubiquinone) 1, alpha/beta subcomplex, 1, 8kDa”.
- ^ a b Wedan, Riley J.; Longenecker, Jacob Z.; Nowinski, Sara M. (January 2024). “Mitochondrial fatty acid synthesis is an emergent central regulator of mammalian oxidative metabolism”. Cell Metabolism. 36 (1): 36–47. doi:10.1016/j.cmet.2023.11.017. PMC 10843818. PMID 38128528.
- ^ Zhang, Shuangxi; Liu, Ruichen; Liu, Quanyu; Liu, Jiankang; Long, Jiangang; Shi, Le (January 2026). “Mitochondrial fatty acid synthesis: The physiopathological role in cellular processes and human diseases”. Genes & Diseases 102034. doi:10.1016/j.gendis.2026.102034.
- ^ a b Norden, Pieter R.; Wedan, Riley J.; Preston, Samuel E. J.; Canfield, Morgan; Graber, Naomi; Longenecker, Jacob Z.; Pentecost, Olivia A.; McLaughlin, Elizabeth; Hart, Madeleine L. (2024-05-10). “Mitochondrial Phosphopantetheinylation is Required for Oxidative Function”. bioRxiv 10.1101/2024.05.09.592977.
- ^ Nowinski, Sara M.; Solmonson, Ashley; Rusin, Scott F.; Maschek, J. Alan; Bensard, Claire L.; Fogarty, Sarah; Jeong, Mi-Young; Lettlova, Sandra; Berg, Jordan A.; Morgan, Jeffrey T.; Ouyang, Yeyun; Naylor, Bradley C.; Paulo, Joao A.; Funai, Katsuhiko; Cox, James E. (2020-08-17). “Mitochondrial fatty acid synthesis coordinates oxidative metabolism in mammalian mitochondria”. eLife. 9 e58041. doi:10.7554/eLife.58041. ISSN 2050-084X. PMC 7470841. PMID 32804083.
- ^ Zong NC, Li H, Li H, Lam MP, Jimenez RC, Kim CS, Deng N, Kim AK, Choi JH, Zelaya I, Liem D, Meyer D, Odeberg J, Fang C, Lu HJ, Xu T, Weiss J, Duan H, Uhlen M, Yates JR, Apweiler R, Ge J, Hermjakob H, Ping P (Oct 2013). “Integration of cardiac proteome biology and medicine by a specialized knowledgebase”. Circulation Research. 113 (9): 1043–53. doi:10.1161/CIRCRESAHA.113.301151. PMC 4076475. PMID 23965338.
- ^ “NDUFAB1 – NADH dehydrogenase [ubiquinone] 1 alpha/beta subcomplex subunit 1”. Cardiac Organellar Protein Atlas Knowledgebase (COPaKB). Archived from the original on 2018-07-20. Retrieved 2018-07-18.
- ^ a b Voet D, Voet JG, Pratt CW (2013). “Chapter 18”. Fundamentals of biochemistry: life at the molecular level (4th ed.). Hoboken, NJ: Wiley. pp. 581–620. ISBN 978-0-470-54784-7.
- ^ a b c d Kim, Dohun; Kesavan, Rushendhiran; Ryu, Kevin; Dey, Trishna; Marckx, Austin; Menezes, Cameron; Praharaj, Prakash P.; Morley, Stewart; Ko, Bookyung; Soflaee, Mona H.; Tom, Harrison J.; Brown, Harrison; Vu, Hieu S.; Tso, Shih-Chia; Brautigam, Chad A. (May 2025). “Mitochondrial NADPH fuels mitochondrial fatty acid synthesis and lipoylation to power oxidative metabolism”. Nature Cell Biology. 27 (5): 790–800. doi:10.1038/s41556-025-01655-4. ISSN 1465-7392. PMC 12331256. PMID 40258949.
- ^ Dibley, Marris G.; Formosa, Luke E.; Lyu, Baobei; Reljic, Boris; McGann, Dylan; Muellner-Wong, Linden; Kraus, Felix; Sharpe, Alice J.; Stroud, David A.; Ryan, Michael T. (January 2020). “The Mitochondrial Acyl-carrier Protein Interaction Network Highlights Important Roles for LYRM Family Members in Complex I and Mitoribosome Assembly”. Molecular & Cellular Proteomics. 19 (1): 65–77. doi:10.1074/mcp.RA119.001784. PMC 6944232. PMID 31666358.
- ^ Dohnálek, Vít; Doležal, Pavel (2024-05-01). “Installation of LYRM proteins in early eukaryotes to regulate the metabolic capacity of the emerging mitochondrion”. Open Biology. 14 (5) 240021. doi:10.1098/rsob.240021. ISSN 2046-2441. PMC 11293456. PMID 38772414.
- ^ Van Vranken, Jonathan G.; Nowinski, Sara M.; Clowers, Katie J.; Jeong, Mi-Young; Ouyang, Yeyun; Berg, Jordan A.; Gygi, Jeremy P.; Gygi, Steven P.; Winge, Dennis R.; Rutter, Jared (August 2018). “ACP Acylation Is an Acetyl-CoA-Dependent Modification Required for Electron Transport Chain Assembly”. Molecular Cell. 71 (4): 567–580. doi:10.1016/j.molcel.2018.06.039. PMC 6104058. PMID 30118679.
- ^ a b Masud, Ali J.; Kastaniotis, Alexander J.; Rahman, M. Tanvir; Autio, Kaija J.; Hiltunen, J. Kalervo (December 2019). “Mitochondrial acyl carrier protein (ACP) at the interface of metabolic state sensing and mitochondrial function”. Biochimica et Biophysica Acta (BBA) – Molecular Cell Research. 1866 (12) 118540. doi:10.1016/j.bbamcr.2019.118540. PMID 31473256.
- ^ Cory, Seth A.; Van Vranken, Jonathan G.; Brignole, Edward J.; Patra, Shachin; Winge, Dennis R.; Drennan, Catherine L.; Rutter, Jared; Barondeau, David P. (2017-07-03). “Structure of human Fe–S assembly subcomplex reveals unexpected cysteine desulfurase architecture and acyl-ACP–ISD11 interactions”. Proceedings of the National Academy of Sciences. 114 (27): E5325–E5334. Bibcode:2017PNAS..114E5325C. doi:10.1073/pnas.1702849114. ISSN 0027-8424. PMC 5502623. PMID 28634302.
- ^ Floyd, Brendan J.; Wilkerson, Emily M.; Veling, Mike T.; Minogue, Catie E.; Xia, Chuanwu; Beebe, Emily T.; Wrobel, Russell L.; Cho, Holly; Kremer, Laura S.; Alston, Charlotte L.; Gromek, Katarzyna A.; Dolan, Brendan K.; Ulbrich, Arne; Stefely, Jonathan A.; Bohl, Sarah L. (2016-08-18). “Mitochondrial Protein Interaction Mapping Identifies Regulators of Respiratory Chain Function”. Molecular Cell. 63 (4): 621–632. doi:10.1016/j.molcel.2016.06.033. ISSN 1097-4164. PMC 4992456. PMID 27499296.
- ^ “NDUFAB1 – NADH dehydrogenase [ubiquinone] 1 alpha/beta subcomplex subunit 1”. UniProt: a hub for protein information. The UniProt Consortium. Retrieved 24 March 2015.
- ^ Otowa T, Maher BS, Aggen SH, McClay JL, van den Oord EJ, Hettema JM (2014-11-12). “Genome-wide and gene-based association studies of anxiety disorders in European and African American samples”. PLOS ONE. 9 (11) e112559. Bibcode:2014PLoSO…9k2559O. doi:10.1371/journal.pone.0112559. PMC 4229211. PMID 25390645.
- ^ Gu Q, Chen XT, Xiao YB, Chen L, Wang XF, Fang J, Chen BC, Hao J (June 2014). “Identification of differently expressed genes and small molecule drugs for Tetralogy of Fallot by bioinformatics strategy”. Pediatric Cardiology. 35 (5): 863–9. doi:10.1007/s00246-014-0868-8. PMID 24463614. S2CID 24162298.