== Background ==
== Background ==
RfaH is a transcription/translation coupling factor found in [[bacteria]], specifically in [[Escherichia coli|”Escherichia coli”]] , that belongs to the NusG family of [[Transcription factor|transcription elongation factor]]<nowiki/>s. As a specialized paralog of NusG, RfaH regulates the expression of long [[Operon|operons]] and has been extensively studied, particularly for its role in activating cell wall biosynthesis, conjugation, and [[Virulence gene|virulence genes]] by inhibiting the [[Rho factor]].<ref name=”:0″>{{Cite journal |last=Bailey |first=Marc J. A. |last2=Hughes |first2=Colin |last3=Koronakis |first3=Vassilis |date=1996 |title=Increased distal gene transcription by the elongation factor RfaH, a specialized homologue of NusG |url=https://onlinelibrary.wiley.com/doi/abs/10.1046/j.1365-2958.1996.d01-1726.x |journal=Molecular Microbiology |language=en |volume=22 |issue=4 |pages=729–737 |doi=10.1046/j.1365-2958.1996.d01-1726.x |issn=1365-2958}}</ref><ref name=”:1″>{{Cite journal|title=Reversible fold-switching controls the functional cycle of the antitermination factor RfaH|url=https://www.nature.com/articles/s41467-019-08567-6|journal=Nature Communications|date=2019-02-11|issn=2041-1723|pmc=6370827|pmid=30742024|volume=10|issue=1|doi=10.1038/s41467-019-08567-6|language=en|first=Philipp Konrad|last=Zuber|first2=Kristian|last2=Schweimer|first3=Paul|last3=Rösch|first4=Irina|last4=Artsimovitch|first5=Stefan H.|last5=Knauer}}</ref> RfaH is characterized as a sequence-specific paralog, which preferentially enhances distal expression within operons that contain specific promoter-proximal ops DNA elements. The ops sequence facilitates the binding of RfaH to elongating [[RNA polymerase]] (RNAP), thereby restricting its functional influence to a limited number of operons within E. coli.<ref name=”:2″>{{Cite journal|title=The β Subunit Gate Loop Is Required for RNA Polymerase Modification by RfaH and NusG|url=https://linkinghub.elsevier.com/retrieve/pii/S1097276511004230|journal=Molecular Cell|date=2011|pmc=3142557|pmid=21777814|pages=253–262|volume=43|issue=2|doi=10.1016/j.molcel.2011.05.026|language=en|first=Anastasia|last=Sevostyanova|first2=Georgiy A.|last2=Belogurov|first3=Rachel A.|last3=Mooney|first4=Robert|last4=Landick|first5=Irina|last5=Artsimovitch}}</ref>
RfaH is a transcription/translation coupling factor found in [[bacteria]], specifically in [[Escherichia coli|”Escherichia coli”]] , that belongs to the NusG family of [[Transcription factor|transcription elongation factor]]<nowiki/>s. As a specialized paralog of NusG, RfaH regulates the expression of long [[Operon|operons]] and has been extensively studied, particularly for its role in activating cell wall biosynthesis, conjugation, and [[Virulence gene|virulence genes]] by inhibiting the [[Rho factor]].<ref name=”:0″>{{Cite journal |last=Bailey |first=Marc J. A. |last2=Hughes |first2=Colin |last3=Koronakis |first3=Vassilis |date=1996 |title=Increased distal gene transcription by the elongation factor RfaH, a specialized homologue of NusG |url=https://onlinelibrary.wiley.com/doi/abs/10.1046/j.1365-2958.1996.d01-1726.x |journal=Molecular Microbiology |language=en |volume=22 |issue=4 |pages=729–737 |doi=10.1046/j.1365-2958.1996.d01-1726.x |issn=1365-2958}}</ref><ref name=”:1″>{{Cite journal|title=Reversible fold-switching controls the functional cycle of the antitermination factor RfaH|url=https://www.nature.com/articles/s41467-019-08567-6|journal=Nature Communications|date=2019-02-11|issn=2041-1723|pmc=6370827|pmid=30742024|volume=10|issue=1|doi=10.1038/s41467-019-08567-6|language=en|first=Philipp Konrad|last=Zuber|first2=Kristian|last2=Schweimer|first3=Paul|last3=Rösch|first4=Irina|last4=Artsimovitch|first5=Stefan H.|last5=Knauer}}</ref> RfaH is characterized as a sequence-specific paralog, which preferentially enhances distal expression within operons that contain specific promoter-proximal ops DNA elements. The ops sequence facilitates the binding of RfaH to elongating [[RNA polymerase]] (RNAP), thereby restricting its functional influence to a limited number of operons within E. coli.<ref name=”:2″>{{Cite journal|title=The β Subunit Gate Loop Is Required for RNA Polymerase Modification by RfaH and NusG|url=https://linkinghub.elsevier.com/retrieve/pii/S1097276511004230|journal=Molecular Cell|date=2011|pmc=3142557|pmid=21777814|pages=253–262|volume=43|issue=2|doi=10.1016/j.molcel.2011.05.026|language=en|first=Anastasia|last=Sevostyanova|first2=Georgiy A.|last2=Belogurov|first3=Rachel A.|last3=Mooney|first4=Robert|last4=Landick|first5=Irina|last5=Artsimovitch}}</ref>
== RfaH Domain architecture (NTD and CTD) ==
== RfaH Domain architecture (NTD and CTD) ==
RfaH has two main [[Protein domain|structural domain protein]], N-terminal domain (NTD) and C-terminal domain (CTD) with which are connected by a flexible linker.<ref name=”:3″>{{Cite journal|title=An α Helix to β Barrel Domain Switch Transforms the Transcription Factor RfaH into a Translation Factor|url=https://linkinghub.elsevier.com/retrieve/pii/S0092867412007702|journal=Cell|date=July 2012|pmc=3430373|pmid=22817892|pages=291–303|volume=150|issue=2|doi=10.1016/j.cell.2012.05.042|language=en|first=Björn M.|last=Burmann|first2=Stefan H.|last2=Knauer|first3=Anastasia|last3=Sevostyanova|first4=Kristian|last4=Schweimer|first5=Rachel A.|last5=Mooney|first6=Robert|last6=Landick|first7=Irina|last7=Artsimovitch|first8=Paul|last8=Rösch}}</ref><ref name=”:0″ /><ref>{{Cite journal|title=Impact of N‐Terminal Domain Conformation and Domain Interactions on RfaH Fold Switching|url=https://onlinelibrary.wiley.com/doi/10.1002/prot.26755|journal=Proteins: Structure, Function, and Bioinformatics|date=March 2025|issn=0887-3585|pages=608–619|volume=93|issue=3|doi=10.1002/prot.26755|language=en|first=Bahman|last=Seifi|first2=Stefan|last2=Wallin}}</ref> The NTDs exhibit mixed α/β topology that comprised of a four-stranded antiparallel [[Beta sheet|β sheet]] surrounded by two and one [[Alpha helix|α helices]] on each sides.<ref name=”:1″ /> The CTD has two long antiparallel α helices that form a [[coiled coil]]. RfaH CTD is closely linked to the NTD and takes on an all-α fold in the free state. When NTD bound to [[RNA polymerase|RNAP]], the domains separate and the CTD transform into an all-β fold, while the NTD remains mostly intact. This existence of CTD in two distinct folded states makes RfaH a classic metamorphic or “transformer” protein. <ref name=”:1″ /><ref>{{Cite journal|title=Structural Basis for Converting a General Transcription Factor into an Operon-Specific Virulence Regulator|url=https://linkinghub.elsevier.com/retrieve/pii/S1097276507001219|journal=Molecular Cell|date=April 2007|pages=117–129|volume=26|issue=1|doi=10.1016/j.molcel.2007.02.021|language=en|first=Georgiy A.|last=Belogurov|first2=Marina N.|last2=Vassylyeva|first3=Vladimir|last3=Svetlov|first4=Sergiy|last4=Klyuyev|first5=Nick V.|last5=Grishin|first6=Dmitry G.|last6=Vassylyev|first7=Irina|last7=Artsimovitch}}</ref>
RfaH has two main [[Protein domain|structural domain protein]], N-terminal domain (NTD) and C-terminal domain (CTD) which are connected by a flexible linker.<ref name=”:3″>{{Cite journal|title=An α Helix to β Barrel Domain Switch Transforms the Transcription Factor RfaH into a Translation Factor|url=https://linkinghub.elsevier.com/retrieve/pii/S0092867412007702|journal=Cell|date=July 2012|pmc=3430373|pmid=22817892|pages=291–303|volume=150|issue=2|doi=10.1016/j.cell.2012.05.042|language=en|first=Björn M.|last=Burmann|first2=Stefan H.|last2=Knauer|first3=Anastasia|last3=Sevostyanova|first4=Kristian|last4=Schweimer|first5=Rachel A.|last5=Mooney|first6=Robert|last6=Landick|first7=Irina|last7=Artsimovitch|first8=Paul|last8=Rösch}}</ref><ref name=”:0″ /><ref>{{Cite journal|title=Impact of N‐Terminal Domain Conformation and Domain Interactions on RfaH Fold Switching|url=https://onlinelibrary.wiley.com/doi/10.1002/prot.26755|journal=Proteins: Structure, Function, and Bioinformatics|date=March 2025|issn=0887-3585|pages=608–619|volume=93|issue=3|doi=10.1002/prot.26755|language=en|first=Bahman|last=Seifi|first2=Stefan|last2=Wallin}}</ref> The NTDs exhibit mixed α/β topology that comprised of a four-stranded antiparallel [[Beta sheet|β sheet]] surrounded by two and one [[Alpha helix|α helices]] on each sides.<ref name=”:1″ /> The CTD has two long antiparallel α helices that form a [[coiled coil]]. RfaH CTD is closely linked to the NTD and takes on an all-α fold in the free state. When NTD bound to [[RNA polymerase|RNAP]], the domains separate and the CTD transform into an all-β fold, while the NTD remains mostly intact. This existence of CTD in two distinct folded states makes RfaH a classic metamorphic or protein. <ref name=”:1″ /><ref>{{Cite journal|title=Structural Basis for Converting a General Transcription Factor into an Operon-Specific Virulence Regulator|url=https://linkinghub.elsevier.com/retrieve/pii/S1097276507001219|journal=Molecular Cell|date=April 2007|pages=117–129|volume=26|issue=1|doi=10.1016/j.molcel.2007.02.021|language=en|first=Georgiy A.|last=Belogurov|first2=Marina N.|last2=Vassylyeva|first3=Vladimir|last3=Svetlov|first4=Sergiy|last4=Klyuyev|first5=Nick V.|last5=Grishin|first6=Dmitry G.|last6=Vassylyev|first7=Irina|last7=Artsimovitch}}</ref>
=== Structural State of RfaH CTD ===
=== Structural State of RfaH CTD ===
<gallery>
File:Structures of RfaH from E. coli.tif|Rfa CTD transformation from an α-hairpin to a β-barrel. <ref name=”:1″ />
Upon recruitment of RfaH to the paused transcription elongation complex at the ops hairpin, the NTD engages RNAP and the CTD is forcibly displaced. Once released, the CTD undergoes a dramatic refolding event into a five-stranded [[Beta barrel|β-barrel]]<ref>{{Cite journal|title=The β Subunit Gate Loop Is Required for RNA Polymerase Modification by RfaH and NusG|url=https://www.cell.com/molecular-cell/abstract/S1097-2765(11)00423-0|journal=Molecular Cell|date=2011-07-22|issn=1097-2765|pmc=3142557|pmid=21777814|pages=253–262|volume=43|issue=2|doi=10.1016/j.molcel.2011.05.026|language=English|first=Anastasia|last=Sevostyanova|first2=Georgiy A.|last2=Belogurov|first3=Rachel A.|last3=Mooney|first4=Robert|last4=Landick|first5=Irina|last5=Artsimovitch}}</ref>, topologically equivalent to the NusG-CTD. NMR analyses of isolated CTD<ref name=”:3″ />, along with computational modeling, show that the thermodynamically preferred state in the absence of interaction between NTD and CTD, is the [[Beta barrel|β-barrel]].<ref>{{Cite journal|title=Multifunnel Landscape of the Fold-Switching Protein RfaH-CTD|url=https://doi.org/10.1021/acs.jpcb.7b11352|journal=The Journal of Physical Chemistry B|date=2018-02-08|issn=1520-6106|pmc=5823028|pmid=29323497|pages=1600–1607|volume=122|issue=5|doi=10.1021/acs.jpcb.7b11352|first=Nathan A.|last=Bernhardt|first2=Ulrich H. E.|last2=Hansmann}}</ref><ref>{{Cite journal|title=Molecular Dynamics Investigations of the α-Helix to β-Barrel Conformational Transformation in the RfaH Transcription Factor|url=https://doi.org/10.1021/jp502193v|journal=The Journal of Physical Chemistry B|date=2014-05-15|issn=1520-6106|pages=5101–5108|volume=118|issue=19|doi=10.1021/jp502193v|first=Jeevan B.|last=GC|first2=Yuba R.|last2=Bhandari|first3=Bernard S.|last3=Gerstman|first4=Prem P.|last4=Chapagain}}</ref><ref>{{Cite journal|title=Energy Landscape for Fold-Switching in Regulatory Protein RfaH|url=https://doi.org/10.1021/acs.jctc.8b00912|journal=Journal of Chemical Theory and Computation|date=2019-01-08|issn=1549-9618|pages=731–742|volume=15|issue=1|doi=10.1021/acs.jctc.8b00912|first=Jerelle A.|last=Joseph|first2=Debayan|last2=Chakraborty|first3=David J.|last3=Wales}}</ref><ref name=”:5″>{{Cite journal|title=Mechanism of the All-α to All-β Conformational Transition of RfaH-CTD: Molecular Dynamics Simulation and Markov State Model|url=https://doi.org/10.1021/ct5002279|journal=Journal of Chemical Theory and Computation|date=2014-06-10|issn=1549-9618|pages=2255–2264|volume=10|issue=6|doi=10.1021/ct5002279|first=Shanshan|last=Li|first2=Bing|last2=Xiong|first3=Yuan|last3=Xu|first4=Tao|last4=Lu|first5=Xiaomin|last5=Luo|first6=Cheng|last6=Luo|first7=Jingkang|last7=Shen|first8=Kaixian|last8=Chen|first9=Mingyue|last9=Zheng|first10=Hualiang|last10=Jiang}}</ref> In this β-form, the CTD gains a new functional surface that is required for downstream interactions, particularly with [[ribosomal protein]] S10 where it facilitates translation when canonical ribosome recruitment elements is absent.<ref name=”:4″ /><ref name=”:5″ />
Upon recruitment of RfaH to the paused transcription elongation complex at the ops hairpin, the NTD engages RNAP and the CTD is forcibly displaced. Once released, the CTD undergoes a dramatic refolding event into a five-stranded [[Beta barrel|β-barrel]]<ref>{{Cite journal|title=The β Subunit Gate Loop Is Required for RNA Polymerase Modification by RfaH and NusG|url=https://www.cell.com/molecular-cell/abstract/S1097-2765(11)00423-0|journal=Molecular Cell|date=2011-07-22|issn=1097-2765|pmc=3142557|pmid=21777814|pages=253–262|volume=43|issue=2|doi=10.1016/j.molcel.2011.05.026|language=English|first=Anastasia|last=Sevostyanova|first2=Georgiy A.|last2=Belogurov|first3=Rachel A.|last3=Mooney|first4=Robert|last4=Landick|first5=Irina|last5=Artsimovitch}}</ref>, topologically equivalent to the NusG-CTD. NMR analyses of isolated CTD<ref name=”:3″ />, along with computational modeling, show that the thermodynamically preferred state in the absence of interaction between NTD and CTD, is the [[Beta barrel|β-barrel]].<ref>{{Cite journal|title=Multifunnel Landscape of the Fold-Switching Protein RfaH-CTD|url=https://doi.org/10.1021/acs.jpcb.7b11352|journal=The Journal of Physical Chemistry B|date=2018-02-08|issn=1520-6106|pmc=5823028|pmid=29323497|pages=1600–1607|volume=122|issue=5|doi=10.1021/acs.jpcb.7b11352|first=Nathan A.|last=Bernhardt|first2=Ulrich H. E.|last2=Hansmann}}</ref><ref>{{Cite journal|title=Molecular Dynamics Investigations of the α-Helix to β-Barrel Conformational Transformation in the RfaH Transcription Factor|url=https://doi.org/10.1021/jp502193v|journal=The Journal of Physical Chemistry B|date=2014-05-15|issn=1520-6106|pages=5101–5108|volume=118|issue=19|doi=10.1021/jp502193v|first=Jeevan B.|last=GC|first2=Yuba R.|last2=Bhandari|first3=Bernard S.|last3=Gerstman|first4=Prem P.|last4=Chapagain}}</ref><ref>{{Cite journal|title=Energy Landscape for Fold-Switching in Regulatory Protein RfaH|url=https://doi.org/10.1021/acs.jctc.8b00912|journal=Journal of Chemical Theory and Computation|date=2019-01-08|issn=1549-9618|pages=731–742|volume=15|issue=1|doi=10.1021/acs.jctc.8b00912|first=Jerelle A.|last=Joseph|first2=Debayan|last2=Chakraborty|first3=David J.|last3=Wales}}</ref><ref name=”:5″>{{Cite journal|title=Mechanism of the All-α to All-β Conformational Transition of RfaH-CTD: Molecular Dynamics Simulation and Markov State Model|url=https://doi.org/10.1021/ct5002279|journal=Journal of Chemical Theory and Computation|date=2014-06-10|issn=1549-9618|pages=2255–2264|volume=10|issue=6|doi=10.1021/ct5002279|first=Shanshan|last=Li|first2=Bing|last2=Xiong|first3=Yuan|last3=Xu|first4=Tao|last4=Lu|first5=Xiaomin|last5=Luo|first6=Cheng|last6=Luo|first7=Jingkang|last7=Shen|first8=Kaixian|last8=Chen|first9=Mingyue|last9=Zheng|first10=Hualiang|last10=Jiang}}</ref> In this β-form, the CTD gains a new functional surface that is required for downstream interactions, particularly with [[ribosomal protein]] S10 where it facilitates translation when canonical ribosome recruitment elements is absent.<ref name=”:4″ /><ref name=”:5″ />
== Mechanism of CTD Metamorphosis ==
== Mechanism of CTD Metamorphosis ==
<gallery>
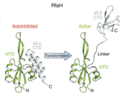
File:Reversible switching between the autoinhibited and active states of RfaH.webp|The process by which the CTD switches from an α-hairpin to a β-barrel—essentially its “metamorphosis”—is driven by activation of RfaH at the ops-paused transcription complex. When the NTD recognizes the ops sequence, the normally tight interface between the NTD and CTD loosens. This is the key event that “releases” the CTD from its inhibited state. Once released, the α-helical hairpin is no longer stable on its own, so it begins to unfold from an all-α helical hairpin to an all-β strand. This new state enables RfaH to interact with translation machinery.<ref name=”:1″ />
”’Domain dissociation:”’ RfaH NTD binds to the RNAP leading to the release of CTD.<ref name=”:6″ />
”’Domain dissociation:”’ RfaH NTD binds to the RNAP leading to the release of CTD.<ref name=”:6″ />
”’Refolding of CTD”’ (”’α → β)”’: CTD dissociation triggers its metamorphic fold switching from all-α helical hairpin to an all-β five-stranded β-barrel thereby activating RfaH.<ref name=”:2″ />
”’Refolding of CTD”’ (”’α → β)”’: CTD dissociation triggers its metamorphic fold switching from all-α helical hairpin to an all-β five-stranded β-barrel thereby activating RfaH.<ref name=”:2″ />
”’Downstream processes:”’ The activated RfaH:opsEC complex then moves downstream with RNAP, where the β-CTD recruits ribosomal protein S10, enabling RfaH to assemble a transcription–translation expressome and promote processive transcription of long operons.
”’Downstream processes:”’ The activated RfaH:opsEC complex then moves downstream with RNAP, where the β-CTD recruits ribosomal protein S10, enabling RfaH to assemble a transcription–translation expressome and promote processive transcription of long operons.
”’Termination:”’ Transcription is terminated when RNAP arrives at the operon terminal. This leads to the dissociation of RfaH dissociates from the complex, allowing the CTD to refold back into the α-helical state and rebind the NTD<ref name=”:1″ />.
”’Termination:”’ Transcription is terminated when RNAP arrives at the operon terminal. This leads to the dissociation of RfaH from the complex, allowing the CTD to refold back into the α-helical state and rebind the NTD<ref name=”:1″ />.
== Biological Importance and Implication ==
== Biological Importance and Implication ==
# ”’Transcriptional regulation”’: RfaH helps to prevent premature termination of specific gene transcription by binding to RNA polymerase. RfaH is a primary transcript elongation regulator through a process known as antitermination.<ref name=”:3″ /> <ref name=”:6″ />
# ”’Transcriptional regulation”’: RfaH helps to prevent premature termination of specific gene transcription by binding to . RfaH is a primary transcript elongation regulator through a process known as antitermination.<ref name=”:3″ /> <ref name=”:6″ />
# ”’Translation factor”’: The refolded RfaH-CTD interacts with the 30S subunit and initiator tRNA, enabling a translation initiation complex to scan nascent mRNA for a start signal, despite the absence of a strong Shine-Dalgarno element. If this recruitment and contact with ribosome is maintained, it enhances translation, reduces Rho-dependent termination by shielding mRNA and possibly preventing ribosome pausing and release.<ref name=”:3″ />
# ”’Translation factor”’: The refolded RfaH-CTD interacts with the 30S subunit and initiator tRNA, enabling a translation initiation complex to scan nascent mRNA for a start signal. If this recruitment and contact with ribosome is maintained, it enhances translation, reduces Rho-dependent termination by shielding mRNA and possibly preventing ribosome pausing and release.<ref name=”:3″ />
# ”’Polarity suppression”’: It suppresses the “polarity” effect that can cause the transcription of distal genes in an operon to be significantly reduced compared to proximal genes.<ref>{{Cite journal|title=Increased distal gene transcription by the elongation factor RfaH, a specialized homologue of NusG|url=https://onlinelibrary.wiley.com/doi/10.1046/j.1365-2958.1996.d01-1726.x|journal=Molecular Microbiology|date=November 1996|issn=0950-382X|pages=729–737|volume=22|issue=4|doi=10.1046/j.1365-2958.1996.d01-1726.x|language=en|first=Marc J. A.|last=Bailey|first2=Colin|last2=Hughes|first3=Vassilis|last3=Koronakis}}</ref>
# ”’Polarity suppression”’: It suppresses the “polarity” effect that can cause the transcription of distal genes in an operon to be significantly reduced compared to proximal genes.<ref>{{Cite journal|title=Increased distal gene transcription by the elongation factor RfaH, a specialized homologue of NusG|url=https://onlinelibrary.wiley.com/doi/10.1046/j.1365-2958.1996.d01-1726.x|journal=Molecular Microbiology|date=November 1996|issn=0950-382X|pages=729–737|volume=22|issue=4|doi=10.1046/j.1365-2958.1996.d01-1726.x|language=en|first=Marc J. A.|last=Bailey|first2=Colin|last2=Hughes|first3=Vassilis|last3=Koronakis}}</ref>
# ”’Enzyme stabilization”’: RfaH stabilizes the RNA polymerase complex, ensuring it can continue moving along the DNA and finish transcribing target long operons.<ref name=”:2″ /><ref name=”:0″ />
# ”’Enzyme stabilization”’: RfaH stabilizes the complex, ensuring it can continue moving along the DNA and finish transcribing target long operons.<ref name=”:2″ /><ref name=”:0″ />
# ”’Regulation of virulence and adaptation”’: RfaH is pivotal for bacterial virulence, host colonization, and survival in hostile environments by regulating the expression of genes involved in cell surface structures like lipopolysaccharide.<ref name=”:7″>{{Cite journal|title=Interdomain contacts control folding of transcription factor RfaH|url=https://academic.oup.com/nar/article/41/22/10077/2436698|journal=Nucleic Acids Research|date=2013-12-01|issn=1362-4962|pmc=3905879|pmid=23990324|pages=10077–10085|volume=41|issue=22|doi=10.1093/nar/gkt779|language=en|first=Sushil Kumar|last=Tomar|first2=Stefan H.|last2=Knauer|first3=Monali|last3=NandyMazumdar|first4=Paul|last4=Rösch|first5=Irina|last5=Artsimovitch}}</ref><ref>{{Cite journal|title=RfaH is essential for virulence and adaptive responses in Yersinia pseudotuberculosis infection|url=https://journals.asm.org/doi/10.1128/mbio.02122-25|journal=mBio|date=2025-09-29|pmc=12607645|pmid=41020597|pages=e02122–25|volume=16|issue=11|doi=10.1128/mbio.02122-25|first=Joram Kiriga|last=Waititu|first2=Kristina|last2=Nilsson|first3=Gerald|last3=Larrouy-Maumus|first4=Tiago R. D.|last4=Costa|first5=Kemal|last5=Avican}}</ref>
# ”’Regulation of virulence and adaptation”’: RfaH is pivotal for bacterial virulence, host colonization, and survival in hostile environments by regulating the expression of genes involved in cell surface structures like lipopolysaccharide.<ref name=”:7″>{{Cite journal|title=Interdomain contacts control folding of transcription factor RfaH|url=https://academic.oup.com/nar/article/41/22/10077/2436698|journal=Nucleic Acids Research|date=2013-12-01|issn=1362-4962|pmc=3905879|pmid=23990324|pages=10077–10085|volume=41|issue=22|doi=10.1093/nar/gkt779|language=en|first=Sushil Kumar|last=Tomar|first2=Stefan H.|last2=Knauer|first3=Monali|last3=NandyMazumdar|first4=Paul|last4=Rösch|first5=Irina|last5=Artsimovitch}}</ref><ref>{{Cite journal|title=RfaH is essential for virulence and adaptive responses in Yersinia pseudotuberculosis infection|url=https://journals.asm.org/doi/10.1128/mbio.02122-25|journal=mBio|date=2025-09-29|pmc=12607645|pmid=41020597|pages=e02122–25|volume=16|issue=11|doi=10.1128/mbio.02122-25|first=Joram Kiriga|last=Waititu|first2=Kristina|last2=Nilsson|first3=Gerald|last3=Larrouy-Maumus|first4=Tiago R. D.|last4=Costa|first5=Kemal|last5=Avican}}</ref>
# ”’Fold-switching”’: RfaH CTD can switch between different three-dimensional shapes (all α helices in free state → β-barrel in activated state) to perform different functions, a property that allows it to be a key regulator in conditions requiring both transcription and translation.<ref name=”:7″ />
# ”’Fold-switching”’: RfaH CTD can switch between different three-dimensional shapes (all α helices in free state → β-barrel in activated state) to perform different functions, a property that allows it to be a key regulator in conditions requiring both transcription and translation.<ref name=”:7″ />
== See also ==
== See also ==
RfaH is a transcription/translation coupling factor found in bacteria, specifically in Escherichia coli , that belongs to the NusG family of transcription elongation factors. As a specialized paralog of NusG, RfaH regulates the expression of long operons and has been extensively studied, particularly for its role in activating cell wall biosynthesis, conjugation, and virulence genes by inhibiting the Rho factor.[1][2] RfaH is characterized as a sequence-specific paralog, which preferentially enhances distal expression within operons that contain specific promoter-proximal ops DNA elements. The ops sequence facilitates the binding of RfaH to elongating RNA polymerase (RNAP), thereby restricting its functional influence to a limited number of operons within E. coli.[3]
RfaH Domain architecture (NTD and CTD)
[edit]
RfaH has two main structural domain protein, N-terminal domain (NTD) and C-terminal domain (CTD) which are connected by a flexible linker.[4][1][5] The NTDs exhibit mixed α/β topology that comprised of a four-stranded antiparallel β sheet surrounded by two and one α helices on each sides.[2] The CTD has two long antiparallel α helices that form a coiled coil. RfaH CTD is closely linked to the NTD and takes on an all-α fold in the free state. When NTD bound to RNAP, the domains separate and the CTD transform into an all-β fold, while the NTD remains mostly intact. This existence of CTD in two distinct folded states makes RfaH a classic metamorphic or transformer protein. [2][6]
Structural State of RfaH CTD
[edit]
-
Rfa CTD transformation from an α-hairpin to a β-barrel. [2]
The CTD of RfaH populates two completely different folded states depending on whether the protein is in its autoinhibited or active form.[4] In the closed, autoinhibited state, the CTD forms a compact two-helical α-hairpin arranged in an antiparallel topology that tightly packs against the NTD.[7] This arrangement masks the RNAP-binding interface on the NTD and keeps RfaH inactive until it encounters its specific DNA recruitment signal on the transcription complex. Structural studies show that this α-helical fold is not intrinsically stable on its own; instead, it is stabilized by extensive NTD–CTD interactions, including buried hydrophobic patches and a well-defined interface where both domains move as a single rigid body[4][2][8][9]
Upon recruitment of RfaH to the paused transcription elongation complex at the ops hairpin, the NTD engages RNAP and the CTD is forcibly displaced. Once released, the CTD undergoes a dramatic refolding event into a five-stranded β-barrel[10], topologically equivalent to the NusG-CTD. NMR analyses of isolated CTD[4], along with computational modeling, show that the thermodynamically preferred state in the absence of interaction between NTD and CTD, is the β-barrel.[11][12][13][7] In this β-form, the CTD gains a new functional surface that is required for downstream interactions, particularly with ribosomal protein S10 where it facilitates translation when canonical ribosome recruitment elements is absent.[8][7]
-
The process by which the CTD switches from an α-hairpin to a β-barrel—essentially its “metamorphosis”—is driven by activation of RfaH at the ops-paused transcription complex. When the NTD recognizes the ops sequence, the normally tight interface between the NTD and CTD loosens. This is the key event that “releases” the CTD from its inhibited state. Once released, the α-helical hairpin is no longer stable on its own, so it begins to unfold from an all-α helical hairpin to an all-β strand. This new state enables RfaH to interact with translation machinery.[2]
Activation: RNAP encounters an operon polarity suppressor (ops) sequence thereby exposing a DNA hairpin (non-template DNA strand) which serve as signal for the recruitment of RfaH to the paused elongation complex (EC).[14]
Domain dissociation: RfaH NTD binds to the RNAP leading to the release of CTD.[14]
Refolding of CTD (α → β): CTD dissociation triggers its metamorphic fold switching from all-α helical hairpin to an all-β five-stranded β-barrel thereby activating RfaH.[3]
Downstream processes: The activated RfaH: opsEC complex then moves downstream with RNAP, where the β-CTD recruits ribosomal protein S10, enabling RfaH to assemble a transcription–translation expressome and promote processive transcription of long operons.
Termination: Transcription is terminated when RNAP arrives at the operon terminal. This leads to the dissociation of RfaH from the complex, allowing the CTD to refold back into the α-helical state and rebind the NTD[2].
Biological Importance and Implication
[edit]
- Transcriptional regulation: RfaH helps to prevent premature termination of specific gene transcription by binding to RNAP. RfaH is a primary transcript elongation regulator through a process known as antitermination.[4] [14]
- Translation factor: The refolded RfaH-CTD interacts with the 30S subunit and initiator tRNA, enabling a translation initiation complex to scan nascent mRNA for a start signal. If this recruitment and contact with ribosome is maintained, it enhances translation, reduces Rho-dependent termination by shielding mRNA and possibly preventing ribosome pausing and release.[4]
- Polarity suppression: It suppresses the “polarity” effect that can cause the transcription of distal genes in an operon to be significantly reduced compared to proximal genes.[15]
- Enzyme stabilization: RfaH stabilizes the RNAP complex, ensuring it can continue moving along the DNA and finish transcribing target long operons.[3][1]
- Regulation of virulence and adaptation: RfaH is pivotal for bacterial virulence, host colonization, and survival in hostile environments by regulating the expression of genes involved in cell surface structures like lipopolysaccharide.[16][17]
- Fold-switching: RfaH CTD can switch between different three-dimensional shapes (all α helices in free state → β-barrel in activated state) to perform different functions, a property that allows it to be a key regulator in conditions requiring both transcription and translation.[16]
- ^ a b c Bailey, Marc J. A.; Hughes, Colin; Koronakis, Vassilis (1996). “Increased distal gene transcription by the elongation factor RfaH, a specialized homologue of NusG”. Molecular Microbiology. 22 (4): 729–737. doi:10.1046/j.1365-2958.1996.d01-1726.x. ISSN 1365-2958.
- ^ a b c d e f g Zuber, Philipp Konrad; Schweimer, Kristian; Rösch, Paul; Artsimovitch, Irina; Knauer, Stefan H. (2019-02-11). “Reversible fold-switching controls the functional cycle of the antitermination factor RfaH”. Nature Communications. 10 (1). doi:10.1038/s41467-019-08567-6. ISSN 2041-1723. PMC 6370827. PMID 30742024.
- ^ a b c Sevostyanova, Anastasia; Belogurov, Georgiy A.; Mooney, Rachel A.; Landick, Robert; Artsimovitch, Irina (2011). “The β Subunit Gate Loop Is Required for RNA Polymerase Modification by RfaH and NusG”. Molecular Cell. 43 (2): 253–262. doi:10.1016/j.molcel.2011.05.026. PMC 3142557. PMID 21777814. ;
- ^ a b c d e f Burmann, Björn M.; Knauer, Stefan H.; Sevostyanova, Anastasia; Schweimer, Kristian; Mooney, Rachel A.; Landick, Robert; Artsimovitch, Irina; Rösch, Paul (July 2012). “An α Helix to β Barrel Domain Switch Transforms the Transcription Factor RfaH into a Translation Factor”. Cell. 150 (2): 291–303. doi:10.1016/j.cell.2012.05.042. PMC 3430373. PMID 22817892. ; ;
- ^ Seifi, Bahman; Wallin, Stefan (March 2025). “Impact of N‐Terminal Domain Conformation and Domain Interactions on RfaH Fold Switching”. Proteins: Structure, Function, and Bioinformatics. 93 (3): 608–619. doi:10.1002/prot.26755. ISSN 0887-3585.
- ^ Belogurov, Georgiy A.; Vassylyeva, Marina N.; Svetlov, Vladimir; Klyuyev, Sergiy; Grishin, Nick V.; Vassylyev, Dmitry G.; Artsimovitch, Irina (April 2007). “Structural Basis for Converting a General Transcription Factor into an Operon-Specific Virulence Regulator”. Molecular Cell. 26 (1): 117–129. doi:10.1016/j.molcel.2007.02.021.
- ^ a b c Li, Shanshan; Xiong, Bing; Xu, Yuan; Lu, Tao; Luo, Xiaomin; Luo, Cheng; Shen, Jingkang; Chen, Kaixian; Zheng, Mingyue; Jiang, Hualiang (2014-06-10). “Mechanism of the All-α to All-β Conformational Transition of RfaH-CTD: Molecular Dynamics Simulation and Markov State Model”. Journal of Chemical Theory and Computation. 10 (6): 2255–2264. doi:10.1021/ct5002279. ISSN 1549-9618.
- ^ a b Cai, Mengli; Agarwal, Nipanshu; Garrett, Daniel S.; Baber, James; Clore, G. Marius (2024-08-20). “A Transient, Excited Species of the Autoinhibited α-State of the Bacterial Transcription Factor RfaH May Represent an Early Intermediate on the Fold-Switching Pathway”. Biochemistry. 63 (16): 2030–2039. doi:10.1021/acs.biochem.4c00258. ISSN 0006-2960. PMC 11345854. PMID 39088556.
- ^ Galaz-Davison, Pablo; Román, Ernesto A.; Ramírez-Sarmiento, César A. (2021). “The N-terminal domain of RfaH plays an active role in protein fold-switching”. PLoS computational biology. 17 (9): e1008882. doi:10.1371/journal.pcbi.1008882. ISSN 1553-7358. PMC 8454952. PMID 34478435.
{{cite journal}}: CS1 maint: article number as page number (link) CS1 maint: unflagged free DOI (link) - ^ Sevostyanova, Anastasia; Belogurov, Georgiy A.; Mooney, Rachel A.; Landick, Robert; Artsimovitch, Irina (2011-07-22). “The β Subunit Gate Loop Is Required for RNA Polymerase Modification by RfaH and NusG”. Molecular Cell. 43 (2): 253–262. doi:10.1016/j.molcel.2011.05.026. ISSN 1097-2765. PMC 3142557. PMID 21777814.
- ^ Bernhardt, Nathan A.; Hansmann, Ulrich H. E. (2018-02-08). “Multifunnel Landscape of the Fold-Switching Protein RfaH-CTD”. The Journal of Physical Chemistry B. 122 (5): 1600–1607. doi:10.1021/acs.jpcb.7b11352. ISSN 1520-6106. PMC 5823028. PMID 29323497.
- ^ GC, Jeevan B.; Bhandari, Yuba R.; Gerstman, Bernard S.; Chapagain, Prem P. (2014-05-15). “Molecular Dynamics Investigations of the α-Helix to β-Barrel Conformational Transformation in the RfaH Transcription Factor”. The Journal of Physical Chemistry B. 118 (19): 5101–5108. doi:10.1021/jp502193v. ISSN 1520-6106.
- ^ Joseph, Jerelle A.; Chakraborty, Debayan; Wales, David J. (2019-01-08). “Energy Landscape for Fold-Switching in Regulatory Protein RfaH”. Journal of Chemical Theory and Computation. 15 (1): 731–742. doi:10.1021/acs.jctc.8b00912. ISSN 1549-9618.
- ^ a b c Artsimovitch, Irina; Landick, Robert (2002-04-19). “The Transcriptional Regulator RfaH Stimulates RNA Chain Synthesis after Recruitment to Elongation Complexes by the Exposed Nontemplate DNA Strand”. Cell. 109 (2): 193–203. doi:10.1016/S0092-8674(02)00724-9. ISSN 0092-8674. PMID 12007406.
- ^ Bailey, Marc J. A.; Hughes, Colin; Koronakis, Vassilis (November 1996). “Increased distal gene transcription by the elongation factor RfaH, a specialized homologue of NusG”. Molecular Microbiology. 22 (4): 729–737. doi:10.1046/j.1365-2958.1996.d01-1726.x. ISSN 0950-382X.
- ^ a b Tomar, Sushil Kumar; Knauer, Stefan H.; NandyMazumdar, Monali; Rösch, Paul; Artsimovitch, Irina (2013-12-01). “Interdomain contacts control folding of transcription factor RfaH”. Nucleic Acids Research. 41 (22): 10077–10085. doi:10.1093/nar/gkt779. ISSN 1362-4962. PMC 3905879. PMID 23990324.
- ^ Waititu, Joram Kiriga; Nilsson, Kristina; Larrouy-Maumus, Gerald; Costa, Tiago R. D.; Avican, Kemal (2025-09-29). “RfaH is essential for virulence and adaptive responses in Yersinia pseudotuberculosis infection”. mBio. 16 (11): e02122–25. doi:10.1128/mbio.02122-25. PMC 12607645. PMID 41020597.